Introduction: A new AI (Artificial Intelligence) algorithm model has been algorithmically learned to process thousands of chest X-rays and produce corresponding After clinical report, thoracic and lung disease was accurately identified in scanned X-rays with an accuracy comparable to that of a radiologist!
At present, most AI models for diagnosing diseases are trained by machine learning on the basis of human-annotated images. In order for the model to predict a certain pathology with reasonable performance, it must be Provide a large number of expert-labeled training examples for this pathology. This process of obtaining high-quality annotations for certain pathologies is expensive and time-consuming, often resulting in large-scale inefficiencies in clinical workflows.
A new algorithm model called CheXzero is born! It can autonomously “learn” from existing medical examination reports, which are written by researchers in Natural Language Processing (NLP). The related research results were published in Nature Biomedical Engineering under the title “Expert-level detection of pathologies from unannotated chest X-ray images via self-supervised learning” (Figure 1).

Figure 1 Research results (Source: [1])
Studies show that properly trained machine learning models often outperform medical experts in tasks involving medical image interpretation. However, such high levels of performance often require training models using relevant datasets carefully annotated by experts. In the study, the model was shown to perform a pathology classification task with self-supervised models on unannotated chest X-ray images with accuracy comparable to that of radiologists. On an external validation dataset of chest radiographs, the self-supervised model outperformed the fully supervised model in detecting three pathologies (eight in total), and the performance generalized to pathologies for which the model was not explicitly annotated, to a variety of image interpretation tasks and datasets from multiple institutions. The purpose of understanding medical images through AI models can significantly save time and capital costs.
A team of researchers from Harvard Medical School trained the CheXzero model using a publicly available dataset of more than 377,000 chest X-rays and more than 227,000 corresponding clinical reports. The researchers tested the performance of CheXzero using unrelated datasets from two different institutions and another country to see that the model was able to compare images with different terms even when the report included different terms. The corresponding report is matched.
Study found:
01
CheXzero classifies pathology without training on any labeled samples
In the absence of explicit labels, the zero-shot method performed comparable to the radiologist and fully supervised methods for pathologies that were not explicitly labeled during training (Figure 2). Specifically, the self-supervised method is -0.042 points lower than the best-performing fully-supervised model in the CheXpert competition. The model learns features from raw radiology reports as a natural source of supervision. For each pathology, a positive and negative cue (eg “merged” vs “not merged”) was generated. By comparing the model outputs for positive and negative cues, the self-supervised method computes a probability score for pathology, which can be used to classify its presence in chest X-ray images.
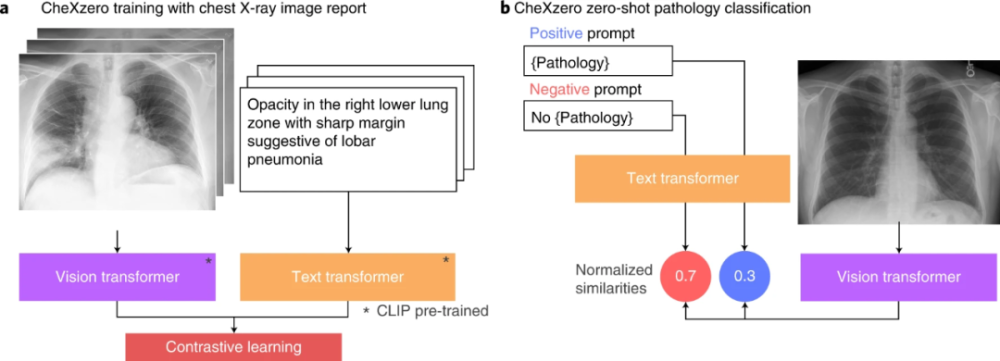
Figure 2 Test process (Source: [1])
Without using explicit labels during training, the self-supervised model outperforms three previous label-efficient methods (MoCo-CXR, MedAug, and ConVIRT) on the CheXpert dataset. MoCo-CXR and MedAug use only chest X-ray images for self-supervision. The self-supervised model achieved these results without using any labels or fine-tuning, showing the model’s ability on zero-shot tasks.
02
CheXzero is significantly better than radiologists in identifying pleural effusions
The F1 score of this model was significantly higher than that of radiologists for pleural effusion, and there were no statistically significant differences for cardiac enlargement, consolidation, and edema. Comparison of ROC curves for self-supervised models with radiologists and test set ground truth. The model outperformed the radiologist when the ROC curve was above the radiologist’s operating point.
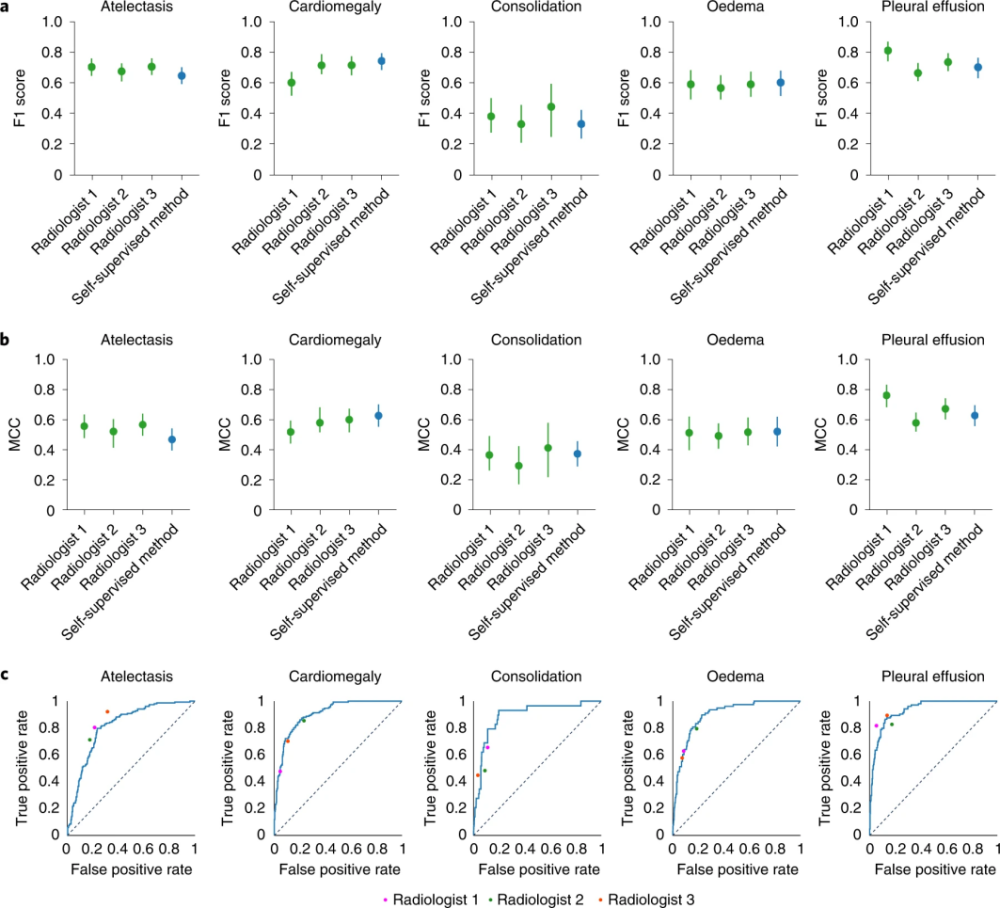
Figure 3 Test process (Source: [1])
This shows that the performance of the self-supervised model is comparable to that of the radiologist, as the performance of the model is comparable to that of the radiologist in the average MCC (Matthews correlation coefficient) of the five CheXpert competing pathologies There was no statistically significant difference between performance on F1.
03
CheXzero has the ability to generalize to datasets in massive amounts of data
Self-supervised methods are able to predict differential diagnosis and radiographic outcomes with high accuracy on datasets collected in different countries than training datasets. This ability to generalize to datasets from disparate distributions has been one of the main challenges in medical AI deployment. A self-supervised model can generalize better because it is able to leverage unstructured textual data, which contains more diverse radiographic information that can be applied to other datasets. Also, of interest in this study is if we use surrogate labels instead of the original clinical findings in PadChest. The results show that the self-supervised approach can generalize well across different data distributions without seeing any clearly labeled pathologies from PadChest during training.
In summary, the self-supervised approach of the new AI algorithm model CheXzero matches radiologist-level performance in a chest X-ray classification task for a variety of pathologies that the model was not explicitly trained to classify. The findings highlight the potential of deep learning models to leverage large amounts of unlabeled data for a wide range of medical image interpretation tasks, which can reduce medical personnel’s reliance on labeled datasets and reduce clinical workflow inefficiencies caused by large-scale labeling.
Pranav Rajpurkar, assistant professor of biomedical informatics at the Blavatnik Institute at Harvard Medical School, led the research project. He said: “We want people to be able to apply the model ‘out of the box’ to other chest X-ray image datasets and disease types that they care about. We are the first to do so and in this The field has effectively demonstrated this. The code for the model has been made public to other researchers, with the hope that it could be applied to CT scans, MRIs and echocardiograms to help detect broader disease in other parts of the body. AI models that require supervised diagnosis can help increase access to healthcare in countries and communities where experts are scarce.”
Christian Leibig, head of machine learning at German startup Vara, said: “Using a richer training signal from the report makes a lot of sense. Vara is using artificial intelligence to detect breast cancer. It can achieve AI detection of the disease. This level of performance is a very big achievement.”
Written by Qiao Weijun
Typesetting|Text Competition
References:
[1]Tiu E, Talius E, Patel P, et al. Expert-level detection of pathologies from unannotated chest X-ray images via self-supervised learning. Nat Biomed Eng. 2022 Sep 15. doi: 10.1038/s41551-022-00936-9. Epub ahead of print. PMID: 36109605.
[2]https:https://www.technologyreview.com/2022/09/15/1059541/ai-medical-notes-teach-itself-spot-disease-chest-x -rays/
[3]https:https://veille-cyber.com/an-ai-used-medical-notes-to-teach-itself/
This article is an original creation of biological exploration. Personal forwarding and sharing are welcome. If any other media or website needs to be reprinted, the source Biological Discovery must be indicated before the text.